Brief Introduction
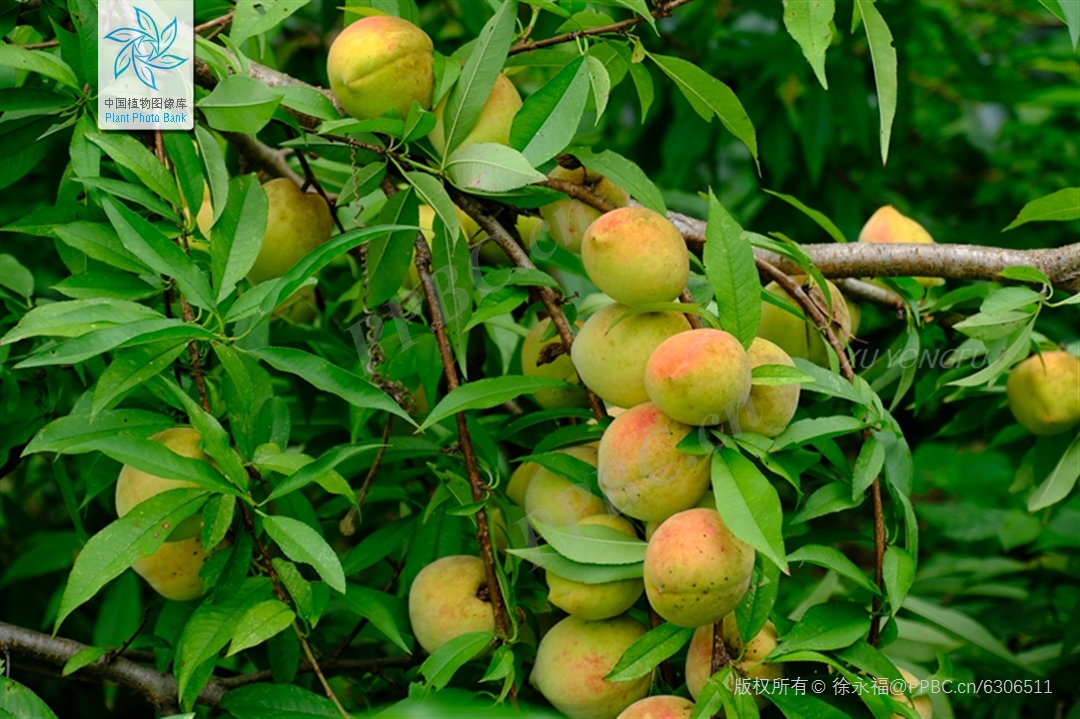
The peach is a globally cultivated economic crop, ranking fourth in global production after apples, pears, and grapes, and also serves as a model species for the Rosaceae family. However, peach breeding is inherently slow and costly due to its perennial nature: trees require several years to flower and fruit, and each plant occupies substantial field space. These limitations highlight the need for early selection tools to improve breeding efficiency.
To address this, we developed PeachDB (http://peachdb.cn/), a comprehensive multi-omics platform for peach that integrates an AI-driven genomic selection model to accelerate breeding and reduce resource input.
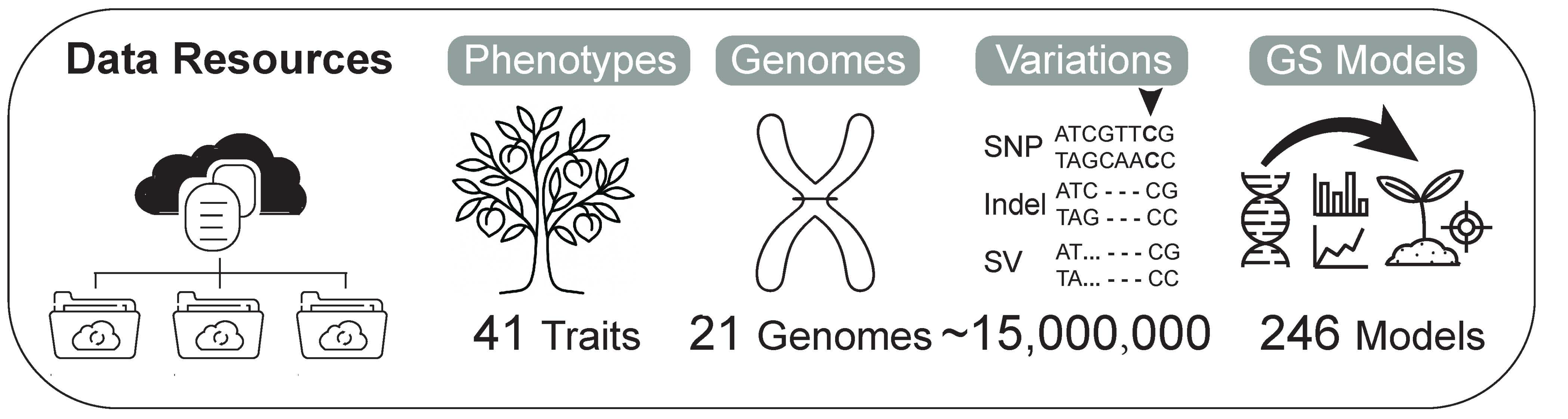
Data resources in PeachDB
We collected data on 41 chemical components and their concentrations relevant to peach fruit quality, including malic acid, citric acid, quinic acid, fructose, glucose, sucrose, and sorbitol.Genomic sequences from 21 peach types, encompassing both wild and cultivated species, specifically Prunus persica, Prunus davidiana, Prunus kansuensis, Prunus ferganensis, and Prunus mira, were analyzed, covering approximately 15,000,000 site variants. In parallel, PeachDB offers 246 Genome Selection (GS) models to facilitate peach breeding programs. Users can download the appropriate model from PeachDB, refer to the accompanying user manual, and input genetic variation data to predict the phenotypic traits of the target population.
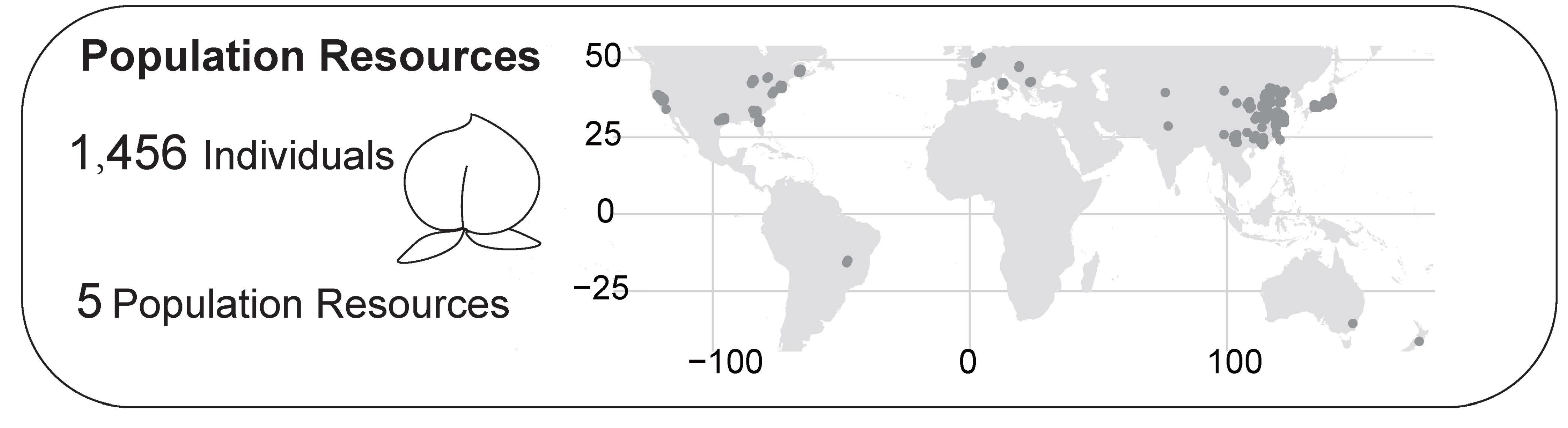
Population resources in PeachDB
Re-sequencing data from 1,456 samples across five global subgroups are compiled.
Available Tools
JBrowse
Browse different types of genomic features such as gene annotaion.
Sequence Blast
Sequence alignment tool by aligning query sequences against a protein/nucleotide reference database.
Phenotype Prediction
Predict phenotypes via Genome Selection models.




